Home > Information > Novel oncogenic mechanism identified through a comprehensive pan-cancer genomic study
-application in cancer precision medicine anticipated-
Novel oncogenic mechanism identified through a comprehensive pan-cancer genomic study
-application in cancer precision medicine anticipated-
April 9, 2020
National Cancer Center Japan
Kyoto University
The Institute of Medical Science, The University Of Tokyo
A research group led by Dr. Keisuke Kataoka at National Cancer Center (President, Dr. Hitoshi Nakagama, Tokyo, Japan), through joint research with Professor Yasushi Okuno of Kyoto University and Professor Satoru Miyano of The Institute of Medical Science, The University Of Tokyo, has completed a comprehensive genomic study of more than 60,000 cancer samples, discovering that multiple mutations within individual oncogenes (hereafter MMs) synergistically promote cancer progression. The results of this research were published on the online edition of the British journal Nature on April 8 (GMT), 2020.
The findings of this study are summarized as follows (Fig. 1):
- It had long been believed that oncogenes gain tumor-promoting functions by acquiring single mutations each individually, but the research team has discovered that MMs are commonly observed in several oncogenes. They were particularly prominent in PIK3CA and EGFR genes, in which 10% of the mutated samples carried MMs respectively. Most of these MMs were located on the same side of the chromosome (in cis).
- Minor (infrequent) mutations were preferentially selected in MMs. Individually, these minor mutations were functionally weak, but synergistically, they exhibited stronger oncogenic potential.
- Samples with PIK3CA MMs showed enhanced downstream pathway activation and higher dependency on the mutated gene itself. They also showed higher sensitivity to specific inhibitors.
The results of this study show that MMs within individual oncogenes serve as a novel genetic mechanism in cancer pathogenesis, also providing an explanation as to why functionally weak minor mutations are accumulated in cancer. In addition, MMs in oncogenes can be exploited as a biomarker for predicting benefits of molecular targeted therapies. Therefore, the application of our findings in cancer precision medicine is anticipated.
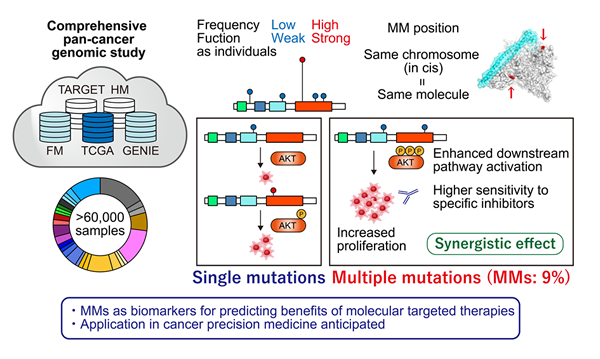
Fig. 1: Summary of the present study
This work was supported by Japan Society for the Promotion of Science (JSPS) KAKENHI (17K19592, 18K06594, and 15H05912), National Cancer Center Research and Development Funds (30-A-1), Ministry of Education, Culture, Sports, Science and Technology (MEXT) as “Priority Issue on Post-K computer" (hp190158 and hp190154), and Foundation for Computational Science (FOCUS) Establishing Supercomputing Center of Excellence.
For further information, please refer to the Japanese version of press release (pdf file).
Details of original paper
Title
Landscape and function of multiple mutations within individual oncogenes
Author
Yuki Saito, Junji Koya, Mitsugu Araki, Yasunori Kogure, Sumito Shingaki, Mariko Tabata, Marni B. McClure, Kota Yoshifuji, Shigeyuki Matsumoto, Yuta Isaka, Hiroko Tanaka, Takanori Kanai, Satoru Miyano, Yuichi Shiraishi, Yasushi Okuno, and Keisuke Kataoka.
Publication
Nature
DOI
10.1038/s41586-020-2175-2
URL
https://www.nature.com/articles/s41586-020-2175-2 (linked at external site)
For media inquiries
National Cancer Center
Office of Public Relations, Strategic Planning Bureau5-1-1 Tsukiji, Chuo-ku, Tokyo 104-0045, Japan
Telephone:+81-3-3542-2511
FAX:+81-3-3542-2545
E-mail:ncc-admin●ncc.go.jp(●を@に置き換えてください)
Kyoto University
Global Communications OfficeYoshidahonmachi, Sakyo-ku, Kyoto 606-8317
Telephone: +81-75-753-5729
FAX: +81-75-753-2094
E-mail:comms●mail2.adm.kyoto-u.ac.jp(●を@に置き換えてください)
The Institute of Medical Science, The University of Tokyo
International Affairs office4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639, Japan
Telephone:+81-3-6409-2027
FAX:+81-3-5449-5496
E-mail:koho●ims.u-tokyo.ac.jp(●を@に置き換えてください)
Files.
- Novel oncogenic mechanism identified through a comprehensive pan-cancer genomic study-application in cancer precision medicine anticipated- (PDF: 889KB)
- 【Japanese version】Novel oncogenic mechanism identified through a comprehensive pan-cancer genomic study-application in cancer precision medicine anticipated- (PDF: 3MB)
