Home > Division of Genome Analysis Platform Development
Division of Genome Analysis Platform Development
Research
Recent advances in high-throughput measurement technology have enabled the comprehensive detection of various types of somatic mutations in cancer genomes. At the same time, information technologies for algorithm and software for accurately and sensitively identifying genuine variants concealed in the enormous amount of noises, deploying these programs into the computing platform such as supercomputer and cloud environment is increasingly becoming important.
We have been developing a practical cancer genome sequencing pipeline (https://genomon -project.github.io/GenomonPagesR/), and the accompanying tools for detecting of acquired mutations using empirical Bayes theories (EBCall, Shiraishi et al., Nucleic Acids Research) and structural variations (GenomonSV, https:// github.com/Genomon-Project/GenomonSV). Through this pipeline, we have contributed many cancer genome projects and identified somatic mutations in splicing machinery in myelodysplastic syndrome, integrated genome, transcriptome, and methylome analysis in kidney cancer (Sato, Yoshizato, Shiraishi et al., Nature Genetics, 2013), various novel pathway mutations in adult T-cell leukemia (Kataoka, Nagata, Kitanaka), Shiraishi et al. , Nature Genetics, 2015) including PD-L1 3’-UTR disruptions (Kataoka, Shiraishi et al., Nature, 2016).
Besides, we have also developed a statistical approach for deriving of mutations signatures (Shiraishi et al., PLoS Genetics 2015), a Bayesian method for identifying of splicing associated variants using genome and transcriptome (Shiraishi et al., Genome Research, 2018) and contributed a large-scale cancer genome analysis project using them (PCAWG Transcriptome Core Group et al., Nature, 2020). Incorporating increasingly evolving information technologies such as machine learning and cloud computing, we would like to develop a genome analysis platform that can contribute to novel discovery by many cancer researchers. We also aim to be deeply involved and discover new insights for ourselves.
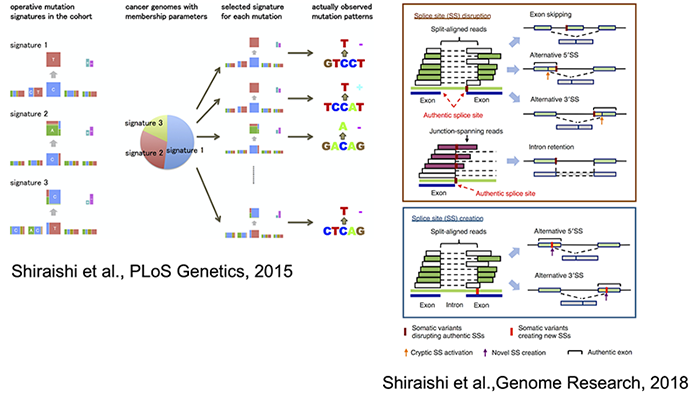
Statistical methods for cancer genomics.

