Home > Organization > Divisions and Independent Research Units > Division of Genome Biology > Research Projects
Research Projects
Genes for personalized therapy
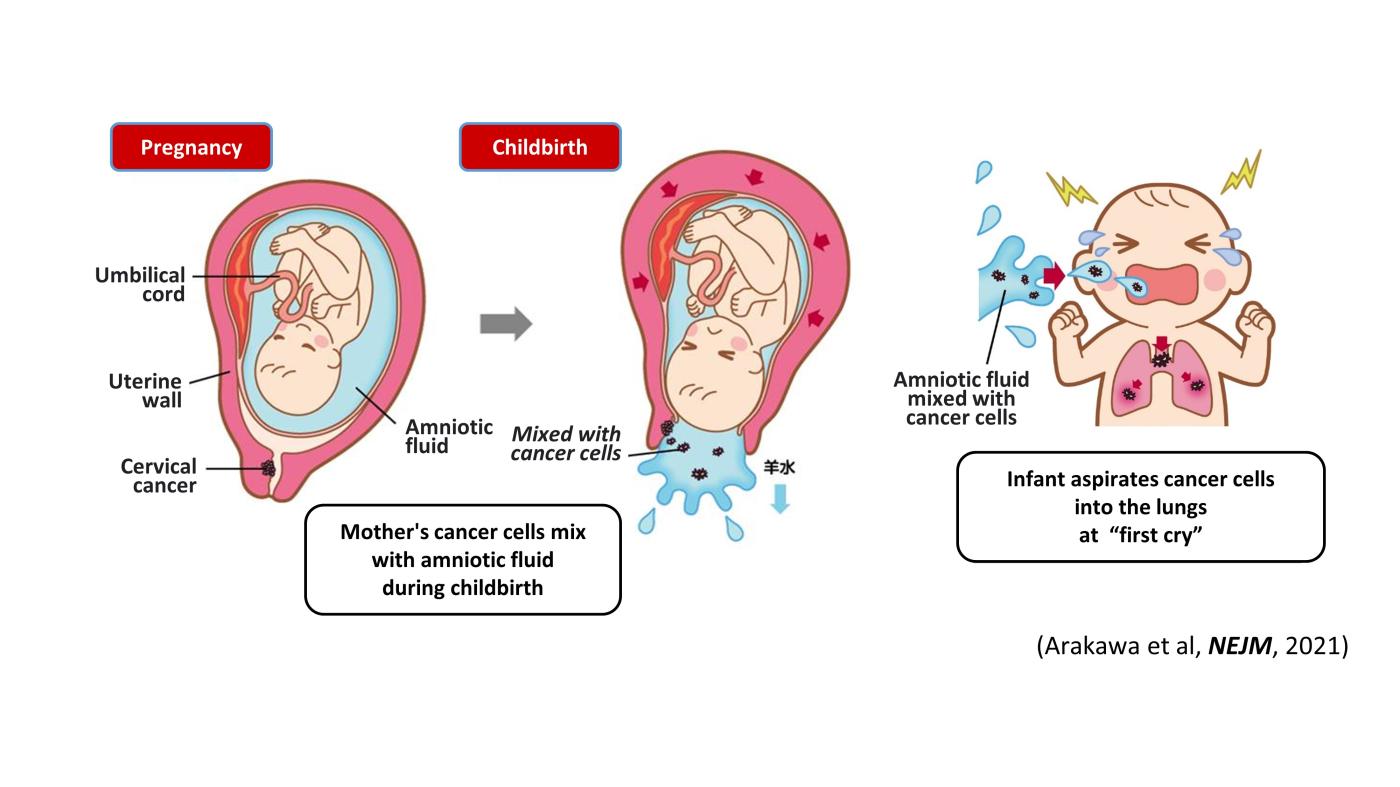
Two cases of pediatric lung cancer (in 23-month-old and 6-year-old boys) resulting from mother-to-infant transmission of uterine cervical tumors were incidentally detected during routine next-generation sequencing of paired samples of tumor and normal tissue. Spontaneous regression of some lesions in the first child and slow growth of the tumor mass in the second child suggested the existence of alloimmune responses against the transmitted tumors. Immune checkpoint inhibitor therapy with nivolumab led to a strong regression of all remaining tumors in the first child.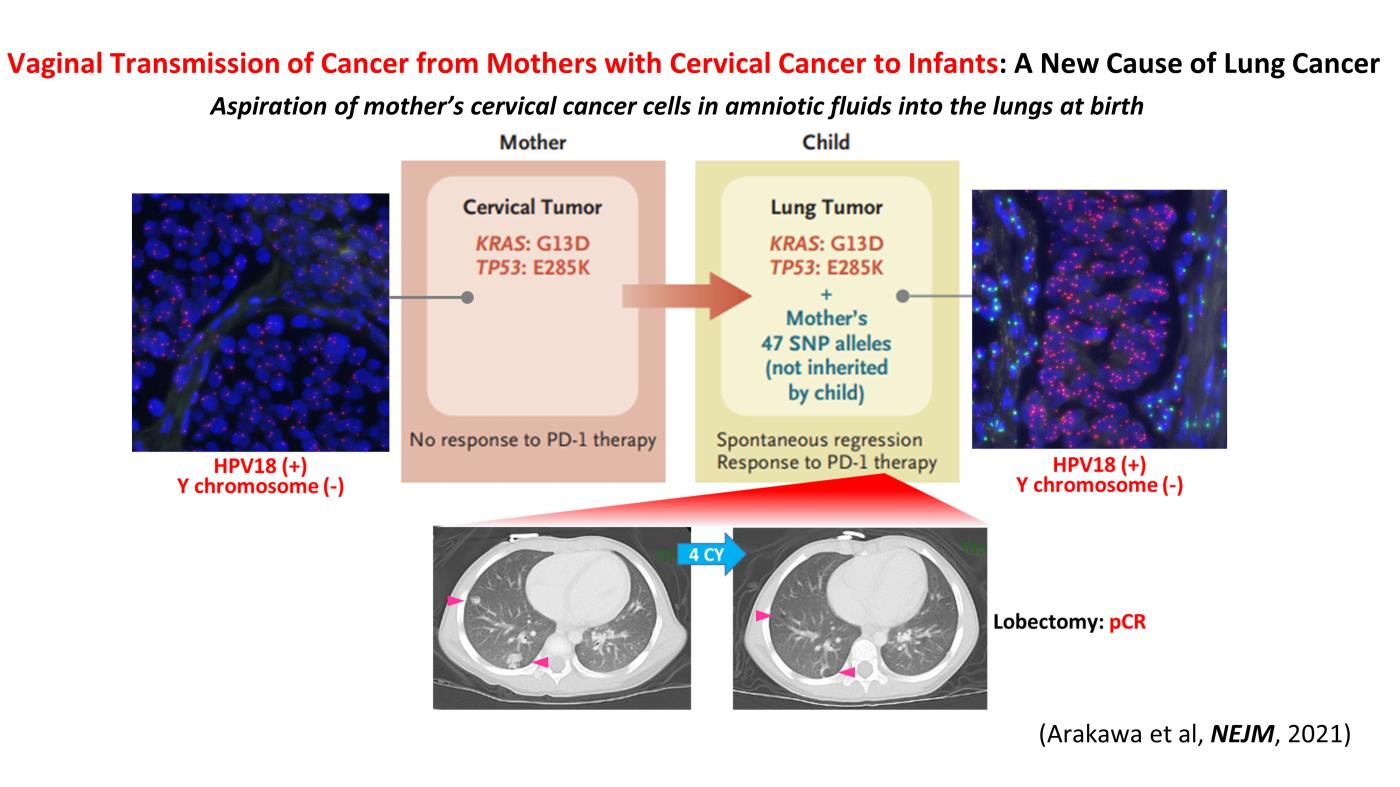
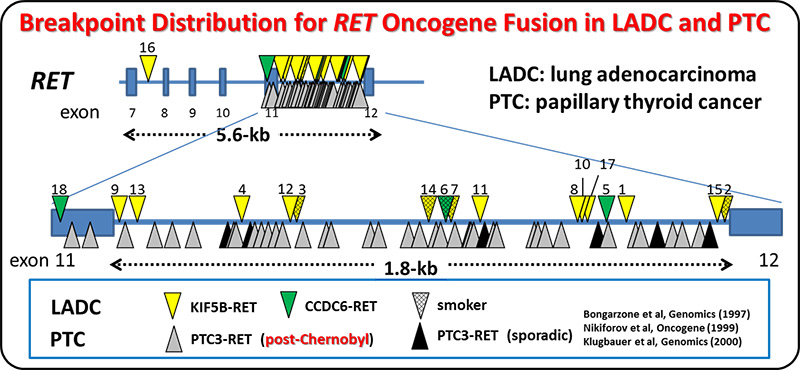
We and others identified the RET fusion gene present in 1-2% of lung adenocarcinomas. LADCs with RET fusion lacked aberrations of other cancer-related genes, supporting the use of monotherapy with RET kinase inhibitor. A Nation-wide genome screening of lung cancer (LC-SCRUM-Japan) involving more than 250 hospitals in Japan, identified 34 RET fusion-positive cases among 1,536 screened ones. A total of 19 patients were enrolled into a phase II investigator-initiated clinical trial (LURET) of a RET TKI, vandetanib. Seventeen cases eligible for efficacy analysis showed a promising result; overall response rate was 53% (90% CI, 31 to 74). By analysing a patient sample enrolled in the LURET trial, we revealed that that missense mutations in the activation loop of RET kinase domain are able to increase
kinase activity and confer drug resistance through allosteric effects. This study demonstrates the significance of “seamless” medical science study, consisting basic, translational, clinical and regulatory science studies, to develop a new therapeutic way.
Analysis of breakpoint junctions for oncogenic RET, ALK and ROS1 fusions revealed that DNA strand breaks formed at non-specific sites within this region trigger RET fusion. Two DNA-repair mechanisms, namely, DNA synthesis-dependent or -independent end-joining pathways were deduced to have contributed to the illegitimate joining of the DNA ends, in the generation of oncogenic fusions.
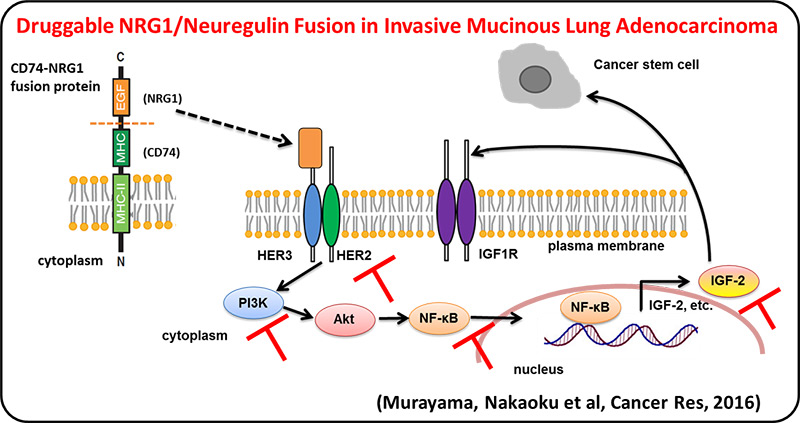
We identified multiple druggable oncogene fusions, including those involving the NRG1 (neuregulin, heregulin), ERBB4, BRAF and RET genes, in invasive mucinous adenocarcinoma (IMA), a malignant type of lung adenocarcinoma. These fusions also represent potentially clinically relevant targets for treatment of IMAs. NRG1 fusion induced NF-kB activity and promoted the IGF2 autocrine/paracrine circuit. Inhibition of ErbB2, PI3K, NF-kB, or IGF2 suppressed NRG1 fusion–induced cancer stem cell–like properties.
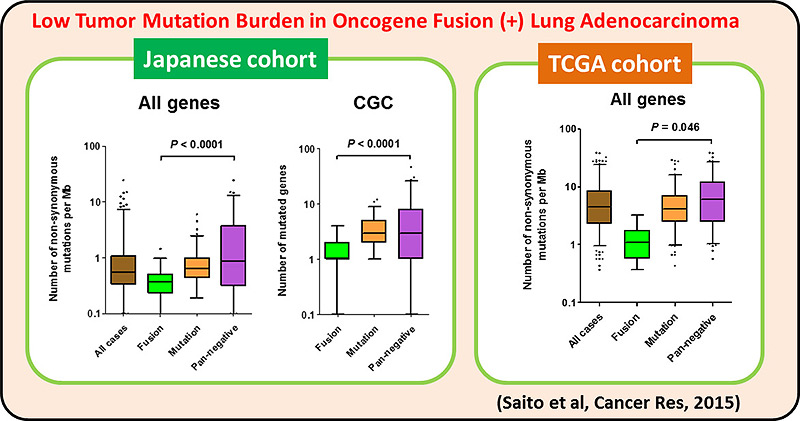
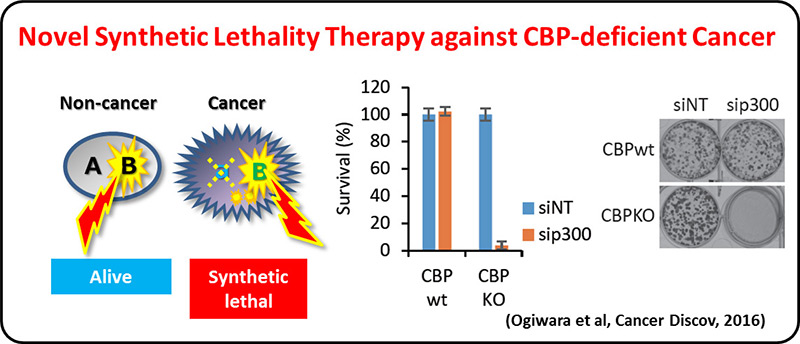
On the other hand, >30% of LADC and most of other types of lung cancers are negative for the oncogene aberrations above, therefore, other therapeutic targets are needed for precision lung cancer medicine. A comprehensive genetic alteration profiling studies of lung cancer genomes, including international ones, revealed inactivation of chromatin-regulating genes, such as SMARCA4/BRG1 and CREBBP, in lung cancer cases negative for oncogene aberrations. We propose a synthetic lethal therapeutic method for chromatin regulator-deficient lung cancers based on inhibition of paralog proteins. A synthetic lethality therapy for CREBBP-deficient small cell lung cancer was proposed; EP300 histone acetyltransferase (functional paralog for CREBBP) is a vulnerability of such cancer, and its functional suppression causes depletion of MYC oncogene product.
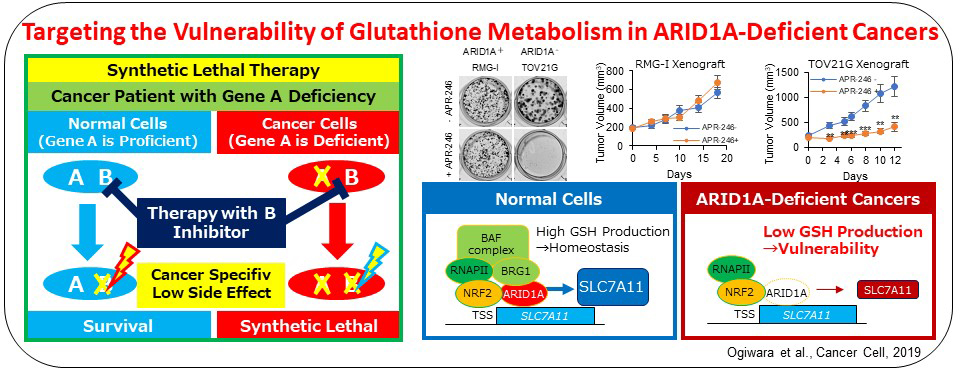
Recently, we propose a synthetic lethal therapeutic method for chromatin regulator ARID1A-deficient cancers. Deleterious ARID1A mutations are frequent in ovarian clear cell (50%) and endometrioid (30%) carcinomas and many other types of cancers, for which there is no effective molecular targeting therapies. We show that cancer cells lacking ARID1A protein expression have a common feature of low SLC7A11 expression caused by impaired ARID1A-mediated transcriptional activation. Such ARID1A-deficient cancer cells were specifically vulnerable to inhibitors of the GSH metabolic pathway, such as APR-246 targeting GSH and buthionine sulfoximine targeting GCLC, due to the low supply of cysteine, a key source of antioxidant GSH, by SLC7A11. Therefore, GCLC-targeting therapy is promising for ovarian and other cancers with ARID1A-deficiency.
Genes for personalized prevention
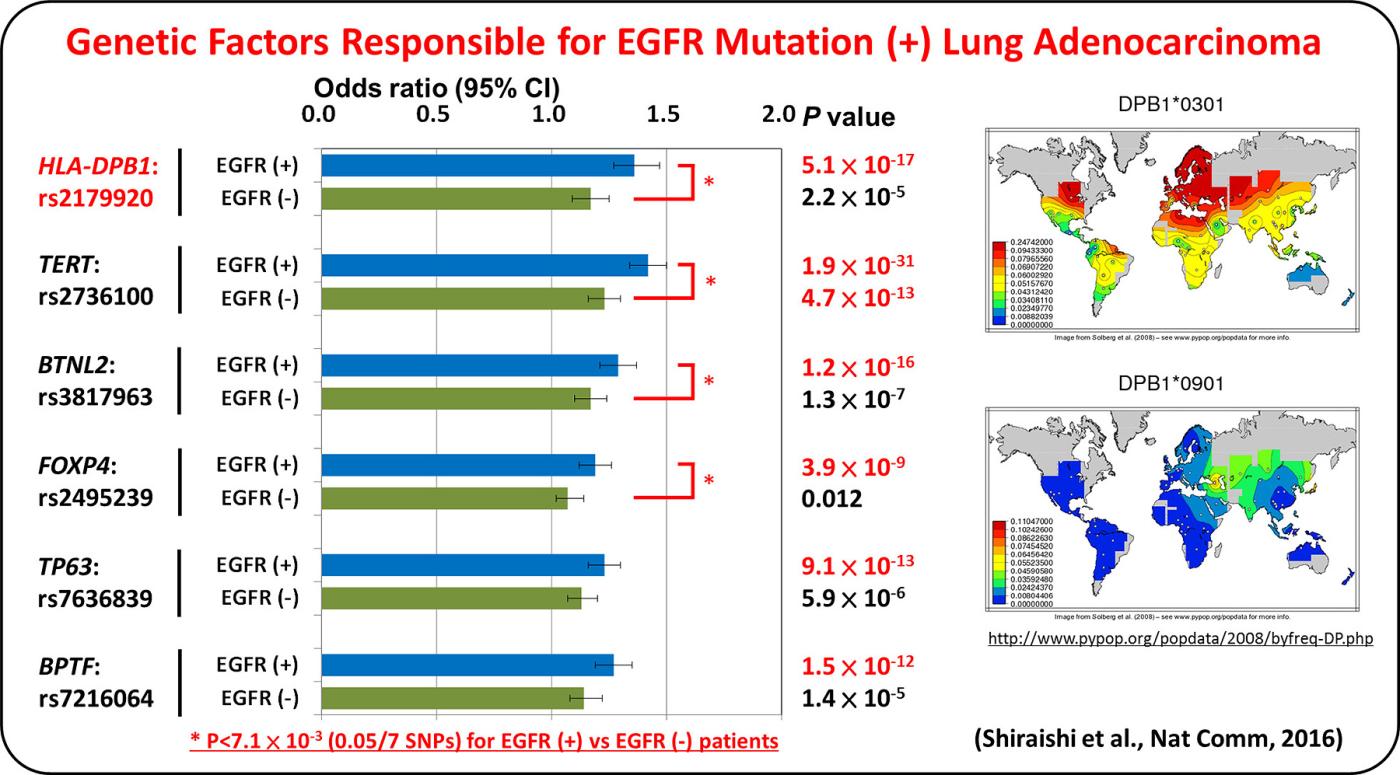
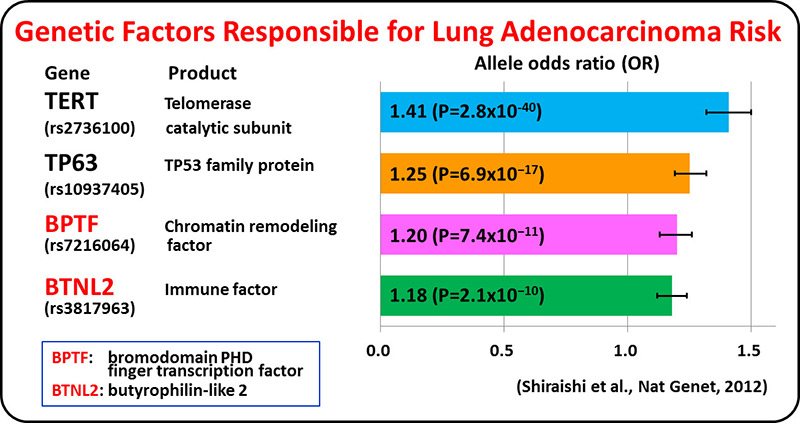
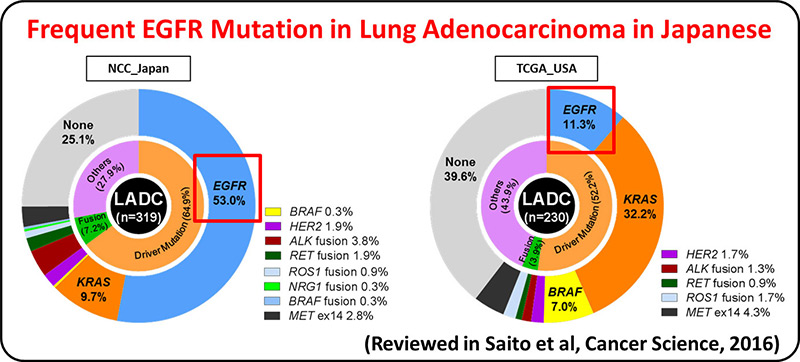
Lung adenocarcinoma driven by somatic EGFR mutations is more prevalent in East Asians (30-50%) than in European/Americans (10-20%), indicating Asian-specific risk factors for lung adenocarcinoma. Genetic factors underlying lung adenocarcinoma risk of Asians are being searched for by performing genome-wide association studies (GWASs), including several international collaborative studies, on SNPs. A genome-wide association study comprising a total of 6,029 individuals with lung adenocarcinoma (cases) and 13,535 controls identified four loci.

GWAS of lung adenocarcinoma with EGFR mutation was performed as a multi-center collaborative study with a Japanese population consisting of 3,173 cases and 15,158 controls. Six loci, including HLA-DPB1, were discovered as risk loci for the disease. The result indicates that lung adenocarcinoma develops in vivo through interaction between somatic mutations and germline variations that modulate immune reaction.