Home > Department of Proteomics
Department of Proteomics
Understanding cancer requires a comprehensive understanding of not only genetic mutations, but also the quantity and activity of proteins that control cell function. We are developing "exploratory proteomics technologies" that use mass spectrometry (*1) and reverse phase protein arrays (*2) for a comprehensive analysis of proteins. Our goal is to get to the heart of cancer (find its weaknesses) at the protein level and link this understanding to the development of treatments. We're also working on developing new ways to analyze proteins, automating the experiments needed for large-scale studies, and creating new techniques for analyzing the data we collect.
*1, Mass Spectrometry (Adachi)
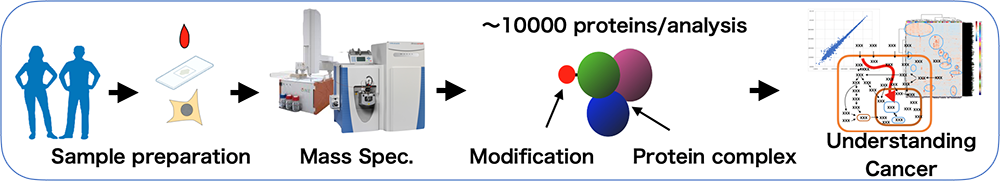
This is a technology that can measure the type and quantity of molecules through precise measurement of molecular size. It allows for the identification and quantification of up to ~10,000 proteins at once. It can also measure protein function and activity through phosphorylation analysis, immunoprecipitation, and proximity labeling etc.
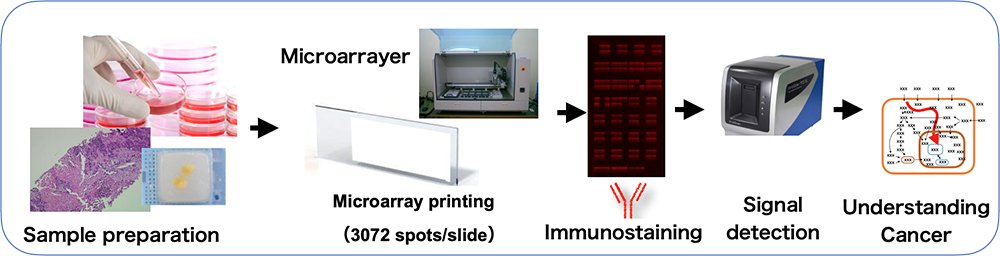
*2, Reverse-Phase Protein Arrays; RPPA (Masuda)
This technology spots lysate derived from cell and tissue samples on a glass slide. Antibodies are then applied to the arrays to quantify the protein of interest in the samples. By using phosphorylation-specific antibodies, it allows us to gain insights into cellular processes, including kinase activity, with high sensitivity.
Project 1: Development of Proteomics Technologies
- Profiling of Cellular Components through Mass Spectrometry
- Functional Dissection of Protein Complexes through Mass Spectrometry
- Quantitative and Functional Analysis of Protein through Reverse-Phase Protein Arrays (RPPA)
Project 2: Proteomics Data Collection Project
- Comprehensive Collection of Cancer Cell Protein Data
- Comprehensive Collection of Phosphorylation Variation Data
- Comprehensive Collection of Antibody Information
Project 3: Experiment Automation Project
- Automation of Sample Processing for Protein Analysis
- Automation of Data Analysis Procedures




