Home > Organization > Divisions and Independent Research Units > Division of Cellular Signaling > Research Projects
Research Projects
Elucidating the Molecular Basis of Cancer and Identifying Therapeutic Targets
Cancer is thought to arise through repeated clonal evolution, driven by the accumulation of random genomic alterations enabled by genomic instability. To develop truly effective therapies for such highly heterogeneous diseases, it is essential to identify the key molecular drivers underlying tumorigenesis in each cancer and to develop drugs that specifically target them. With this in mind, we conduct comprehensive genomic analyses of patient specimens to clarify the mechanisms of carcinogenesis and to discover novel therapeutic targets.
For example, we performed a large-scale, pan-cancer evaluation of the efficacy of targeted protein degraders using patient-derived xenograft (PDX) models, in which tumor tissues obtained from patients are implanted into immunodeficient mice. We further carried out whole-exome sequencing of these PDX models and identified BRCA and ATM mutations as predictive biomarkers of therapeutic response (NPJ Precis Oncol. 8 : 117 , press release )
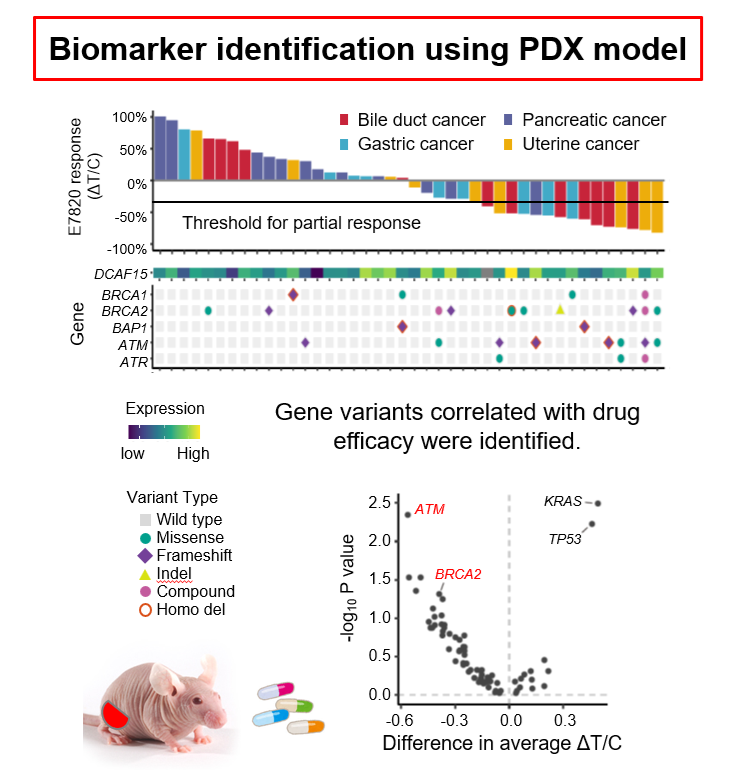
Translational Research in Cancer Genomic Medicine
We are also actively engaged in the clinical implementation of cancer genomic medicine. In genomic medicine, sequence alterations in cancer-related genes are analyzed using next-generation sequencing. To this end, we developed a unique gene panel assay, the TOP panel, which enables the simultaneous extraction and analysis of both DNA and RNA from clinical specimens (Cancer Science. 110, 4 , press release).
Using this platform, it is possible to identify cancer-related genomic alterations across nearly all solid tumors in a single assay. We have also extended this approach to liquid biopsy applications using circulating cell-free DNA and circulating tumor cells in the bloodstream ( Cancer Discov. 13, 8 , press release).
In addition, we successfully developed an original high-throughput method to determine which of the numerous genomic variants identified in this way contribute to tumorigenesis and which are responsible for drug resistance (Sci Transl Med. 9:eaan6556, press release). When we applied this method to more than 1,000 variants of uncertain significance in various cancer-related genes, we found that many of them were indeed involved in cancer development (Nat Commun. 11 : 2573, press release).
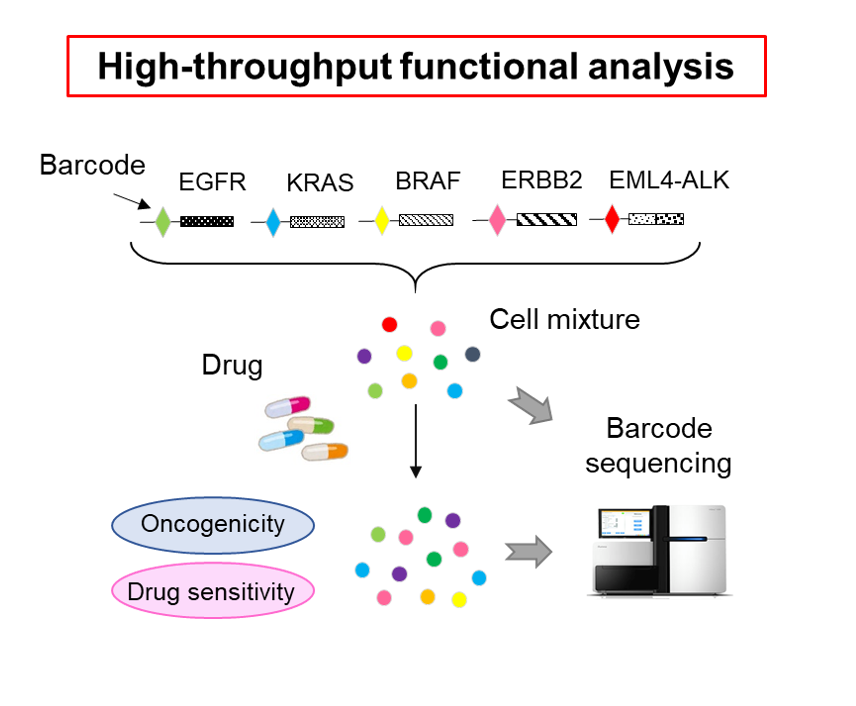
Development of new computational pipeline for cancer research
Moreover, we are establishing new computational pipeline for multi-omic analysis by ourselves to optimize for our research purpose. Our motto is“build what we need”while there are many analytical pipeline available these days. For instance, allele sprcific copy number analytical tool (for in-house use) or FFPE specific mutation error elimination tool named MicroSEC (Commun Biol 4: 1396) and fusion detection tool for genome medicine (Cancer Sci 110: 1464) have been developed.

